DNA Motif Analysis
Unfortunately the large size of the ADAM33 genomic sequence prevents motif discovery based on the DNA sequence with free online tools. MEME and MOTIF Search were able to analyze the mRNA and the results and analysis can be seen below.
Meme Motif Search
Motif Search
mRNA Found Motifs
EGF_1: EGF-like domain signature 1
Prosite: PS00022
Integrin_Beta: Integrins beta chain cysteine-rich domain signature.
Prosite: PS00243
CTCK_1: C-terminal cystine knot signature.
Prosite: PS01185
ANAPHYLATOXIN_1: Anaphylatoxin domain signature.
Prosite: PS01177
4FE4S_FERREDOXIN: 4F2- 4S ferredoxins, iron-sulfur binding region signature.
Prosite: PS00198
VWFC_1: VWFC domain signature.
Prosite:PS01208
2FE2S_FER_1: 2Fe-2S ferredoxins, iron-sulfur binding region signature.
Prosite: PS00197
DEFENSIN: Mammalian defensins signature
Prosite: PS00269
Prosite: PS00022
Integrin_Beta: Integrins beta chain cysteine-rich domain signature.
Prosite: PS00243
CTCK_1: C-terminal cystine knot signature.
Prosite: PS01185
ANAPHYLATOXIN_1: Anaphylatoxin domain signature.
Prosite: PS01177
4FE4S_FERREDOXIN: 4F2- 4S ferredoxins, iron-sulfur binding region signature.
Prosite: PS00198
VWFC_1: VWFC domain signature.
Prosite:PS01208
2FE2S_FER_1: 2Fe-2S ferredoxins, iron-sulfur binding region signature.
Prosite: PS00197
DEFENSIN: Mammalian defensins signature
Prosite: PS00269
Analysis
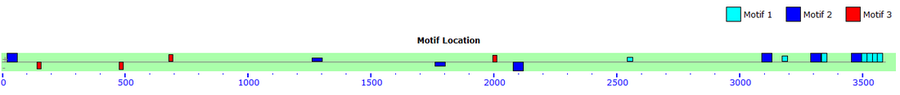
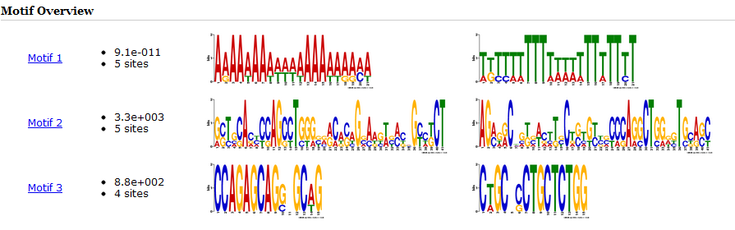
Both online tools were useful for analyzing the mRNA sequence, but in different ways. The MEME Motif was able to identify motifs and displayed them by showing there nucleotide sequence. It was able to identify 3 separate motifs in multiple sites in the mRNA sequence. Motif search provided easier analysis of the predicted function of the mRNA product.
References
- Meme http://meme.sdsc.edu/meme4_6_0/intro.html
- Motif Search http://motif.genome.jp/