Gene Ontology
Gene Ontology is a bioinformatics initiative to standardize gene and gene product attributes across species and databases. The goal is to analyze the function of a protein or gene by identifying common sequences or motifs across all genes. Gene ontology can give a good estimation about physical location in cell and genome, molecular functions, and roles in larger biological networks. Information about the ADAM33 function was obtained using Amigo [1].
Amigo
Amigo Results
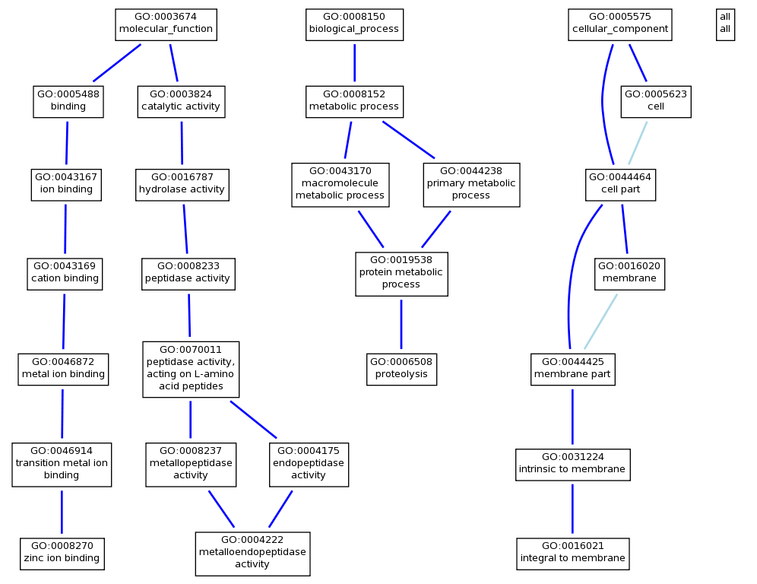
This picture was generated graphically using the data from Amigo. [1] The protein motifs show evidence for the ADAM33 function in proteolysis using Zinc ion binding, and being localized to the membrane. The zinc ion binding, metalloendopeptidase activity, proteolysis, and the protein motif integral to membrane proteins are all predicted using NAS evidence. [2] These results are fairly accurate in representing the function of ADAM33 from the results obtained using Homologene. The crystal structure of the catalytic domain of human ADAM33 also listed four cofactors associated with ADAM33 protein. Calcium ion, zinc ion, chlorine ion, and a N-ACETYL-D-GLUCOSAMINE polysaccharide. [5]
References:
- Amigo http://amigo.geneontology.org/cgi-bin/amigo/go.cgi
- Evidence of Go sites. http://amigo.geneontology.org/cgi-bin/amigo/gp-assoc.cgi?gp=UniProtKB:Q9BZ11&session_id=5206amigo1297980609
- String. http://string-db.org/
- Jie Z, Jin M, Cai Y, Bai C, Shen Y, Yuan Z, Hu Y, Holgate S. The effects of Th2 cytokines on the expression of ADAM33 in allergen-induced chronic airway inflammation. Respir Physiol Neurobiol. 2009 Sep 30;168(3):289-94. Epub 2009 Jul 25.
- Orth, P., et al., and Madison, V., Crystal structre of the catalytic domain of human ADAM33 J.MOL.BIOL. vol:335, pag:129-137 (2004)