Note: Reading through the site and background information will help facilitate understanding of this study proposal.
Study Proposal
Goal: Because the role of ADAM33 in Asthma patheogenesis is not completely understood, a murine model would be extremely useful in understanding more about the disease. This study hopes to produce a mouse line that shows the phenotypical characteristics of Asthma, and to gain clues on the function of ADAM33 in asthma.
Hypothesis: Mice with similar SNP mutations found in Humans will have an altered function, or show phenotypical characteristics similar to asthma patients.
Method: Study mutants with similar polymorphisms found in human linkage analysis studies. The function of ADAM33 will be elucidated in several ways. Expression levels will be monitored by testing with a microarray and real time PCR. The effect of mutations on the expression level of proteins related to ADAM33 or immune system response will be analyzed using a microarray or rtPCR. Novel isoforms or localization problems will be identified by use of a GFP fusion protein. Further details of the methodology and reasoning are found below.
1. Most significant SNPs found in introns or 3’ UTR region.
Eerdeqegh P., Little R. , et al. Association of the ADAM33 gene with asthma and bronchial hyperresponsiveness. Nature. 2002 Jul 25;418(6896):383-4.
|
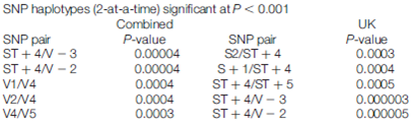
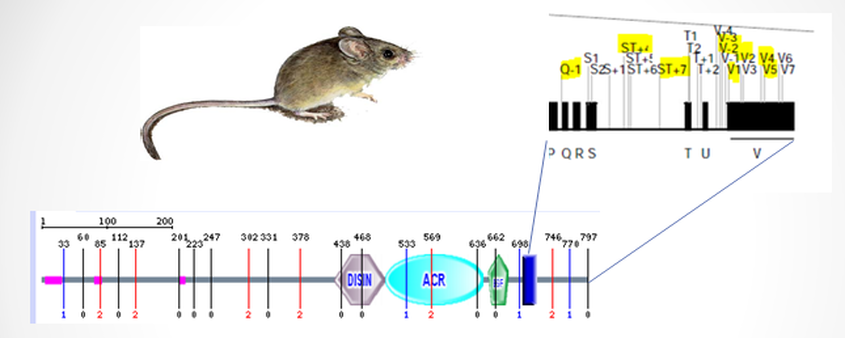
Since the work by Van Eerdewegh [1] and associates identified ADAM33 as an asthma-susceptible gene, many studies have investigated the relationship between ADAM33 polymorphisms and asthmatic conditions. Unfortunately, the sites of mutations have not been consistent across all populations. T1, T2, S2, and V-3 sites were reported in Japanese adult population association studies [5], and the study by
|
Van Eerdewegh [1] showed the ADAM33 polymorphisms (Q-1, S1,
ST+4, V-1, and V4) were associated with childhood asthma in Caucasians. Despite
these differences most studies have shown significant haplotypes and SNP’s have
occurred at the 3’ end of the gene. Many of these mutations are located in the introns between
the trans membrane domain and cytoplasmic domain or in the 3’ UTR region.
Mutations in these regions may affect alternative splicing, splicing efficiency,
mRNA stability, or protein localization. To determine if mutations in these region
can cause asthmatic response or alter function, a mutated ADAM33-GFP fusion
protein will be inserted into ADAM33 asthmatic mice to observe for asthmatic phentoypes.
2. Creating Transgenic Mice
By inserting a mutated gene of ADAM33 back into an ADAM33 knockout mouse, correct insertion of the gene can be easily established. As previous studies have shown, ADAM33 knockouts do not show the phenotypical features of asthma. Correctly mutated mice should show expression of ADAM33, and mice with asthmatic conditions should show an increased pulmonary resistance, IgE count, abnormal level of immune cells, or staining for abnormal lung sections. Analysis of transgenic mice with asthmatic conditions should provide a basis for understanding the role of ADAM33 in asthma patheogenesis.
3. Analyzing the Results
Determining if mutations result in changes in expression or degradation.
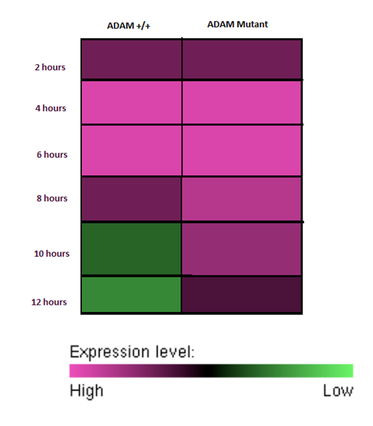
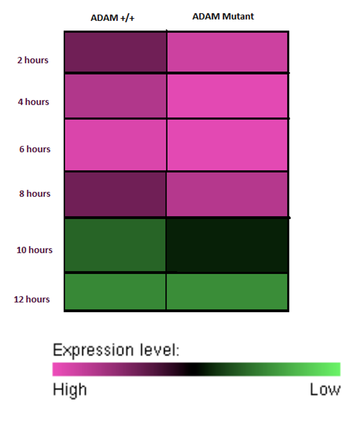
By testing with real time PCR, changes in ADAM33 expression due to mutations can be analyzed. The expression levels of ADAM33 can be compared between mice by comparing to a housekeeper gene. Because ADAM33 expression level has been shown to increase in response to ultra fine carbon particles [3], it is expected that mutations may result in abnormal expression of ADAM33 proteins. Two hypothetical results are seen below.
|
If mRNA degradation is slowed then it is expected the ADAM33 mRNA will increase at relatively the same rate as wild type, but be around for a longer period. |
If mRNA translation is increased it is expected that the ADAM33 mutant mouse will show higher expression levels of ADAM33 proteins at an earlier time point then wildtype mice. |
Determining if mutations change expression level in immune pathway proteins.
|
Using a microarray, a comparison can be made from the ADAM33 wild type and ADAM33 mutant mice to see if additional proteins have either decreased or increased transcription levels. From the background data collected throughout the site, it is expected that the ADAM33 mutant mice will show an increased response in IL-4 proteins and a decreased level of IL-2 proteins. Additionally, other proteins can be analyzed for changes of expression levels. Analyzing results of this microarray will help determine if any additional protein level changes can be attributed to mutations in the ADAM33 gene. |
Determining localization problems with GFP tags.
Question: Do
the mutations effect localization of any of the ADAM33 isoforms,
or result in new isoforms?
As shown in the protein domain page there are three known isoforms of ADAM33 genes. Because many of the SNP and haplotypes positively associated with asthma were located in introns, it is possible that these mutations may result in alternative splicing or effect localization of one of the ADAM33 isoforms.
Method: Look for unusual locations of ADAM33 proteins using GFP tags. Identify isoform changes using 2d gel electrophoresis and identify different isoforms using mass spectrometry.
Expected Result: Isoforms or localizations may be causing asthmatic phenotypes. If the ADAM33 protein is in a different location, or with a predominantly novel isoform it may change the function of ADAM33 in the asthmatic response pathway. The GFP tags will provide important information in determining the location of most isoforms of the ADAM33 gene and locating novel isoforms.
As shown in the protein domain page there are three known isoforms of ADAM33 genes. Because many of the SNP and haplotypes positively associated with asthma were located in introns, it is possible that these mutations may result in alternative splicing or effect localization of one of the ADAM33 isoforms.
Method: Look for unusual locations of ADAM33 proteins using GFP tags. Identify isoform changes using 2d gel electrophoresis and identify different isoforms using mass spectrometry.
Expected Result: Isoforms or localizations may be causing asthmatic phenotypes. If the ADAM33 protein is in a different location, or with a predominantly novel isoform it may change the function of ADAM33 in the asthmatic response pathway. The GFP tags will provide important information in determining the location of most isoforms of the ADAM33 gene and locating novel isoforms.
4. Confirming the Results
Determining if Asthmatic conditions are a result of over-expression.
Two mice lines can be created that carry different levels of the ADAM33 gene. A mouse line with two copies of the ADAM33 gene can be compared to a mouse line with 4 copies of the ADAM33 to see if an asthmatic phenotype is due solely to the over-expression of the ADAM33 gene. If this is the case, the line with 4 copies of the ADAM33 gene will express more ADAM33 proteins and have a stronger asthmatic response.
5. Looking Towards the Future
Future Direction of Research
If a positive correlation with ADAM33 and asthma can be determined, it is possible that small chemicals that inhibit ADAM33 may work to reduce the asthmatic response in humans. Using Bioassay, two small chemicals were identified that showed inhibition of ADAM33 at different concentration ranges. The results can be seen on the small chemical page. Future studies could explore the possibility of treating asthmatic patients using chemicals that inhibit ADAM33, and the murine model would be an important step in ensuring the safety of the drugs prescribed.
References:
- Eerdeqegh P., Little R. , et al. Association of the ADAM33 gene with asthma and bronchial hyperresponsiveness. Nature. 2002 Jul 25;418(6896):383-4.
- Chen, C. et al. 2006. Mol. Cell. Biol. 26(18):6950-6956
- GEO Profiles: http://www.ncbi.nlm.nih.gov/geoprofiles/
- Dimitris Georgiadis, Athanasios Yiotakis. Specific targeting of metzincin family members with small-molecule inhibitors: Progress toward a multifarious challenge. Bioorganic & Medicinal Chemistry Volume 16, Issue 19, 1 October 2008, Pages 8781-8794