Protein Domains
By using free analysis programs such as Prosite and Smart online, it is possible to get a rough idea of a proteins function. Below I looked at the functional domains of ADAM33 isoform 1, and confirmed the results from an analysis of the crystal structure found on the Protein Database of Japan's website [4]. Additionally information about the three different ADAM33 isoform and post translational modifications are shown below.
Prosite
Analysis of Prosite data:
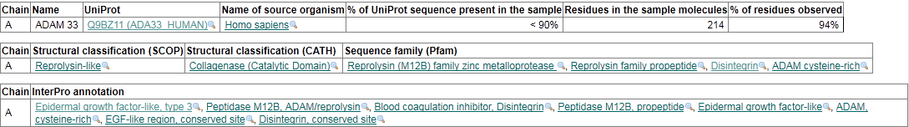
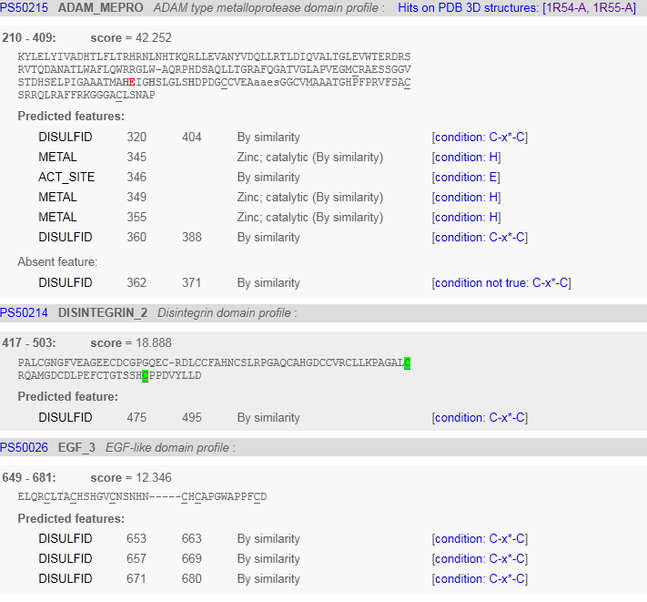
The ADAM_MEPRO is part of the ADAM type metalloprotease domain profile. It is found in between the 210 and 409 amino acids. It is a section of the protein that indicates it is part of the ADAM family of proteins. The ADAM cysteine-rich domain is not found in plant, archaeal, bacterial or viral proteins. ADAMs are transemembrane proteins containing both a disintegrin and a metalloprotease domain. [5] This is a correct analysis, because this is a member of the ADAM family of proteins.
The Disintegrin 2 domain contains an Arg-Gly-Asp sequence that is the recognition site of many adhesion proteins. [6] The RGD sequence of disintegrins is postulated to interact with the glycoprotein IIb-IIIa complex.
The EGF-3 domain signature is a sequence of forty amino-acid residues, known to be present in a large number of mostly animal proteins. A common feature is that these repeats are found in the extracellular domain of membrane-bound proteins or in proteins known to be secreted The EGF
contains disulfide bridges which first bind with high affinity to specific cell-surface receptors then induces their dimerization, which is essential for
activating the tyrosine kinase in the receptor cytoplasmic domain, initiating a signal transdution that results in DNA synthesis and cell proliferation. [5]
Binding to a zinc ion was also correctly predicted by prosite, and was able to identify 3 out of the 4 amino acids involved in binding to a zinc ion. The correct binding sites can be confirmed by the analysis of the crystal structure seen here.
Prosite was unable to predict binding of Calcium ion, chloride ion, and N-ACETYL-D-GLUCOSAMINE from the crystal structure seen here.
SMART
Analysis of Smart
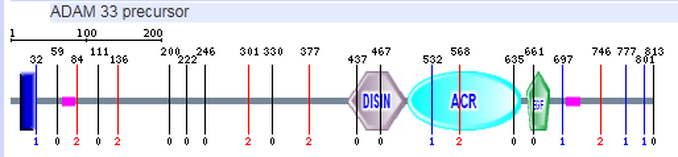
In addition to the DISIN, ACR, and EGF domain that Prosite was able to detect, SMART detected a transmembrane domain (seen in blue) and a region of low complexity (seen in pink). The transmembrane domain was not detected by Prosite or Pfam. Because ADAM33 is a transmembrane protein this is a possible location of the transmembrane region. A Smart analysis of mouse showed the transmembrane region in the area of low complexity shown above. Smart was unable to predict the zinc binding sites that Prosite predicted.
Crystal Structure of the Catylitic domain of ADAM33
A crystal structure was found by searching the Protein Database of Europe and confirmed some of the analysis done by the SMART and Prosite databases. The structure was published by Orth, P., et al., and Madison, V., in
2004 in a paper entitled "Crystal structure of the catalytic domain of
human ADAM33" In addition to domains listed above, four cofactors are associated with the ADAM33 protein. A calcium ion, zinc ion, chlorine ion, and a N-ACETYL-D-GLUCOSAMINE
polysaccharide. [7] Listed below are the domains found using this service.
Information from the crystal structure can be viewed here
Information from the crystal structure can be viewed here
3d Structure of ADAM33 Domains
A 3d model of the Adam 33 protein can be seen here
Protein Isoforms
Three isoforms of ADAM33 can be found from NCBI by following the links below. These isoforms are produced by alternative splicing.
Isoform 1 - Alpha Helix
-mRNA sequence -proprotein sequence
Isoform 2 - Transcript Variant: This variant (2) lacks an in-frame exon in the coding region, as compared to variant 1. It encodes isoform beta, which lacks an internal segment at 636-661, but has the same N- and C-termini as isoform alpha. This structure no longer contains the EGF domain that Isoform 1 contains.
-mRNA sequence -proprotein sequence
Isoform 3 - This form differs from Isoform a1, by missing internal segments at 1-478 and 636-661. There is no experimental conformation available for this isoform. Notes from similarity of mouse isoform.
Isoform 1 - Alpha Helix
-mRNA sequence -proprotein sequence
Isoform 2 - Transcript Variant: This variant (2) lacks an in-frame exon in the coding region, as compared to variant 1. It encodes isoform beta, which lacks an internal segment at 636-661, but has the same N- and C-termini as isoform alpha. This structure no longer contains the EGF domain that Isoform 1 contains.
-mRNA sequence -proprotein sequence
Isoform 3 - This form differs from Isoform a1, by missing internal segments at 1-478 and 636-661. There is no experimental conformation available for this isoform. Notes from similarity of mouse isoform.
Post Translational Modification
- The precursor is cleaved by a furin endopeptidase [9]
- The protein is also phosphorylated at Tyr243 [10][11]
References:
- Kann's Lab: Domain Mapping of Disease Locations. http://bioinf.umbc.edu/DMDM/gene_prot_page.php?id=80332
- Protein Data Base Europe: http://www.ebi.ac.uk/pdbe-srv/view/entry/1r54/ligands.html
- Orth P, Reichert P, Wang W, Prosise WW, Yarosh-Tomaine T, Hammond G, Ingram RN, Xiao L, Mirza UA, Zou J, Strickland C, Taremi SS, Le HV, Madison V.Crystal Structure of the catalytic domain of ADAM 33 http://www.ncbi.nlm.nih.gov/pubmed/14659745?dopt=Abstract
- Protein Database Japan www.pdbj.org
- Prosite http://www.expasy.ch/cgi-bin/prosite/ScanView.cgi?scanfile=852981221711.scan.gz
- Calvete J.J., Schafer W., Soszka T., Lu W.Q., Cook J.J., Jameson B.A., Niewiarowski S. Identification of
the disulfide bond pattern in albolabrin, an RGD-containing peptide from
the venom of Trimeresurus albolabris: significance for the expression
of platelet aggregation inhibitory activity.
- Orth, P., et al., and Madison, V., Crystal structure of the catalytic domain of human ADAM33 J.MOL.BIOL. vol:335, pag:129-137 (2004)
- Smart http://smart.embl-heidelberg.de/smart/
- Van Eerdewegh P, Keith T.P. Association of the ADAM33 gene with asthma and bronchial hyperresponsiveness.
- Rikova K, et al. (2007) Global survey of phosphotyrosine signaling identifies oncogenic kinases in lung cancer. Cell 131, 1190-203
- Phosphorylation Site Page. http://www.phosphosite.org/siteAction.do?id=1005844