What is Asthma?
Asthma has been documented in humans before the creation of the pyramids. The Georg Ebers Papyrus,
found in Egypt in the 1870s, contains prescriptions written in hieroglyphics
for over seven hundred remedies. [9] Despite mankind’s ancient knowledge of
this disease, it was not until the 1960s that the inflammatory component of
asthma was recognized. It is typically triggered by allergens in the air resulting in elevated levels of IgE which cause mast cell activation, chronic airway inflammation, and airway hyper-responsiveness. [8] When an asthma attack occurs, the muscles surrounding the airways become tight and the lining of the air passages swells. [4]
|
Common asthma triggers include:
|
|
The prevalence of asthma has increased significantly since the 1970s. As of 2009, 300 million people were affected worldwide In 2009 asthma caused 250,000 deaths globally.[5]
What is ADAM33?
ADAM33 encodes a disintegrin and metalloprotein-33 protein that participates in the
bronchial remodeling process in asthma [2]. Members of the ADAM family are membrane-anchored proteins that have been implicated in a variety of biological processes involving cell-cell and cell-matrix interactions. [4]
The ADAMs family of transmembrane proteins is part of the zinc protease superfamily. Members of this family are characterized by the presence of metalloprotease, integrin receptor-binding activites, and a cytoplasmic domain that in many family members specifies binding sites for various signal transducing proteins. They have been implicated in the control of membrane fusion, cytokine and growth factor shedding, cell migration, muscle development, fertilization, and cell fate determination.
The ADAMs family of transmembrane proteins is part of the zinc protease superfamily. Members of this family are characterized by the presence of metalloprotease, integrin receptor-binding activites, and a cytoplasmic domain that in many family members specifies binding sites for various signal transducing proteins. They have been implicated in the control of membrane fusion, cytokine and growth factor shedding, cell migration, muscle development, fertilization, and cell fate determination.
How is ADAM33 related to Asthma?
|
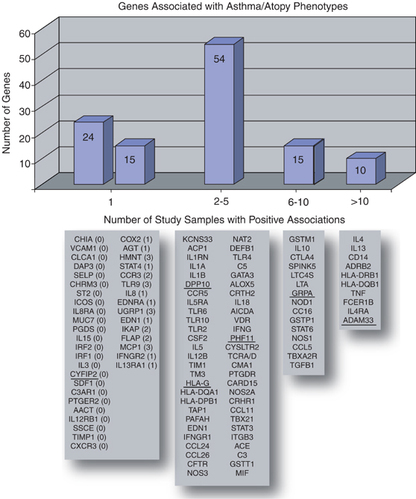
Asthma is a respiratory disease with a remarkably heterogeneous phenotype. The risk, age of onset, and severity of asthma is influenced by numerous biological pathways that interact with environmental exposures. These biological pathways are influenced by multiple genes which additively contribute to asthmatic symptoms [3]. The amount of genes contributing to asthmatic risk may exceed 100, but the individual effect on any one of these genes may vary. It is likely that for any particular phenotype a small number of genes may greatly influence the risk of asthma in an individual. As asthma research continues, a great challenge is finding constellations of genes that interact to determine specific risk for individuals. [3]
Asthma has been associated with over 100 candidate genes in the human genome [6]. Of these candidate genes only 10 genes have been associated in greater then 10 independent studies. Genes confirmed in more then 6 independent studies are generally considered to be true susceptibility genes to asthma related phenotypes. [3] Adam-33 has been confirmed in 15 independent samples. Adam 33 has been strongly associated with objective measures such as bronchial hyper responsiveness and accelerated decline in forced expiratory volume per second. In addition, frequencies for a common SNP in ADAM33 called ST+7 have shown frequencies of 84.9% in the asthmatic population and 79.1% in the control population. Based on an estimated prevalence of 8%, this SNP would potentially contribute to |
around 50 000 excess asthma cases in the UK population [10] The study was conducting using 8 separate population groups (US/UK, Mexican , Puerto Rican, Dutch, US white, German
, Nottingham, Iceland) further validating ADAM33's strong association with asthma. [10]
References
- Bjornsdottir U, Blakey J, Hakonarson H, Halapi E, Kristinsson, Stefansson K, Upmanyu R, S, Wheatley A. Contribution of ADAM33 polymorphisms to the population risk of asthma. I P Hall Thorax 2005;60:274–27
- Candelaria I. Vergaraa, Nathalie Acevedoa, Silvia Jiméneza, Beatriz Martíneza, Dilia Mercadoa, Leonor Gusmãoc, Kathleen C. Barnesd, Luis Caraballo Int A Six-SNP Haplotype of ADAM33 Is Associated with Asthma in a Population of Cartagena, Colombia Arch Allergy Immunol 2010;152:32-40
- Hoffjan S. Ober C. Asthma genetics 2006: the long and winding road to gene discovery. Genes and Immunity (2006) 7, 95–100
- Kaneshiro N. Astham. U.S. National Library of Medicine, National Institutes of health. http://www.ncbi.nlm.nih.gov/pubmedhealth/PMH0001196
- Lemanske R. Global Strategy for Asthma Management and Prevention. Gina Report 2009
- Mahdi Bijanzadeh, Nallur B. Ramachandra, P. A. Mahesh, R. Savitha Mysore, Pradeep Kumar, B. S. Manjunath and B. S. Jayaraj Association of IL-4 and ADAM33 Gene Polymorphisms with Asthma in an Indian Population. Lung, 2010, Volume 188, Number 5, Pages 415-422
- Seals D., Courtneigde S. The ADAMs family of metalloproteases: multidomain proteins with multiple function. Genes and Development. 2003 17: 7-30
- Mdbiosciences http://www.mdbiosciences.com/ova-induced-asthma-whitepaper/
- Hammad, B. N. (2003). Taking our breath away: dendritic cells in the pathogenesis of asthma. Nature Reviews Immunology, 994-1003
- Paul Van Eerdewegh et al. Association of the ADAM33 gene with asthma and bronchial hyperresponsiveness. Nature 418, 426-430
|
|